Transplant Rejection Testing - CAM 329
Description:
Transplant rejection involves an immune response to a transplanted organ. The recipient’s immune system recognizes the donated organ as “foreign,” thereby initiating an immune response as if the transplanted organ was a foreign antigen. This response may cause the transplanted organ to fail.1 Gene expression profiling tests and serum cell-free DNA evaluation are possible ways to monitor organ transplant rejection.2-4
Regulatory Status
Many labs have developed specific tests that they must validate and perform in house. These laboratory-developed tests (LDTs) are regulated by the Centers for Medicare and Medicaid as high-complexity tests under the Clinical Laboratory Improvement Amendments of 1988 (CLIA ’88). LDTs are not approved or cleared by the U. S. Food and Drug Administration; however, FDA clearance or approval is not currently required for clinical use.
AlloMap was approved by the FDA on August 26, 2008, as an In Vitro Diagnostic Multivariate Index assay (IVDMIA) test service, performed in a single laboratory, assessing the gene expression profile of RNA isolated from peripheral blood mononuclear cells (PBMC). AlloMap Testing is intended to aid in the identification of heart transplant recipients with stable allograft function who have a low probability of moderate/severe acute cellular rejection (ACR) at the time of testing in conjunction with standard clinical assessment.5
The Heartsbreath test received approval on February 24, 2004, through the FDA’s humanitarian device exemption. The order is described as being “indicated for use as an aid in the diagnosis of grade 3 heart transplant rejection in patients who have received heart transplants within the preceding year. The heartsbreath test is intended to be used as an adjunct to, and not as a substitute for, endomyocardial biopsy. The use of the device is limited to patients who have had endomyocardial biopsy within the previous month.”21
Policy:
Application of coverage criteria is dependent upon an individual’s benefit coverage at the time of the request.
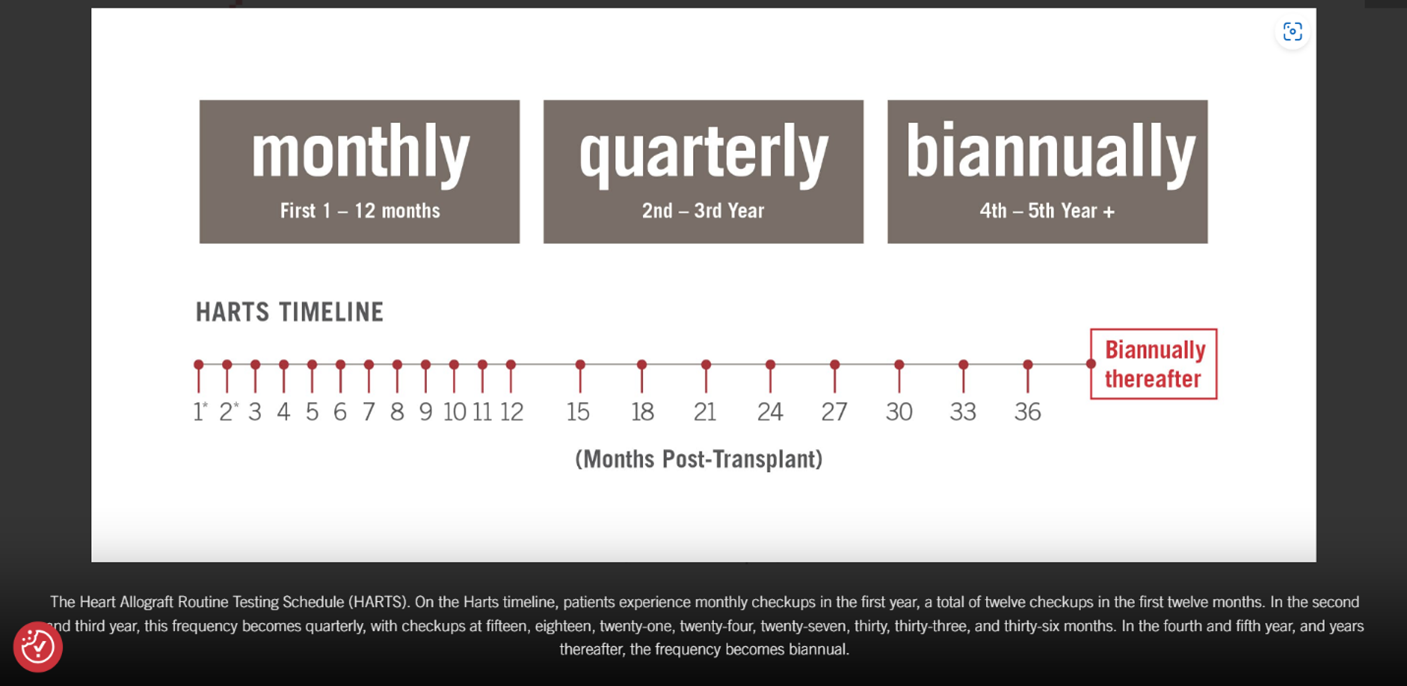
- For heart transplant recipients who are 15 years of age or older and who are 55 or more days post-transplant, the use of AlloMap® is considered MEDICALLY NECESSARY at the following frequency:
- Every month for individuals who are 2 – 12 months post-transplant.
- Every three months for individuals who are 12 – 36 months post-transplant.
- Every six months for individuals who are greater than 36 months post-transplant.
- For heart transplant recipients who are 28 or more days post-transplant, the use of donor-derived cell-free DNA tests (e.g., AlloSure Heart, Prospera™ Heart) is considered MEDICALLY NECESSARY at the following frequency:
- Every month for individuals who are 1 – 12 months post-transplant.
- Every three months for individuals who are 12 – 36 months post-transplant.
- Every six months for individuals who are greater than 36 months post-transplant.
- For single-kidney transplant recipients who are 18 years of age or older and who are at least 14 days post-transplant, the use of donor-derived cell-free DNA tests (e.g., AlloSure Kidney) to assess the probability of allograft rejection is considered MEDICALLY NECESSARY at the following intervals:
- One test per month at one, two, three, and four months post-transplant.
- One test every three months beginning at six months post-transplant.
- For kidney transplant recipients with stable renal function, the use of peripheral blood microarray-based genomic tests that analyze gene expression profiles to rule out kidney transplant rejection (e.g., TruGraf) is considered NOT MEDICALLY NECESSARY.
The following does not meet coverage criteria due to a lack of available published scientific literature confirming that the test(s) is/are required and beneficial for the diagnosis and treatment of an individual’s illness.
- For heart transplant recipients, testing for heart transplant rejection with simultaneous gene expression testing and donor-derived cell-free DNA testing (e.g., HeartCare®) is considered NOT MEDICALLY NECESSARY.
- For any other organ transplant not listed above (e.g., lungs, liver), the use of donor-derived cell-free DNA tests is considered NOT MEDICALLY NECESSARY.
- For any other organ transplant not listed above (e.g., kidneys, lungs, liver), the use of peripheral blood gene expression profiling tests is considered NOT MEDICALLY NECESSARY.
- To assess for rejection and injury in transplanted organs, mRNA expression profiling of biopsied tissue from a transplanted organ (e.g., MMDx Heart, MMDx Kidney) is considered NOT MEDICALLY NECESSARY.
- To assist in the detection of moderate grade 2R/grade 3 heart transplant rejection, the measurement of volatile organic compounds is considered NOT MEDICALLY NECESSARY.
Table of Terminology
| Term |
Definition |
| AASLD |
American Association for the Study of Liver Diseases |
| ACR |
Acute cellular rejection |
| ALAD |
Acute lung allograft dysfunction |
| AMR |
Antibody-mediated rejection |
| AR |
Active rejection |
| AST |
American Society of Transplantation |
| ASTS |
American Society of Transplant Surgeons |
| AUC |
Area under curve |
| BAL |
Bronchoalveolar lavage |
| BL |
Borderline rejection |
| CARGO |
Cardiac Allograft Rejection Gene Expression Observational |
| cfDNA |
Cell-free deoxyribonucleic acid |
| CLAD |
Chronic lung allograft dysfunction |
| CLIA ’88 |
Clinical Laboratory Improvement Amendments Of 1988 |
| CMS |
Centers For Medicare and Medicaid |
| dd-cfDNA |
Donor-derived cell-free deoxyribonucleic acid |
| DNA |
Deoxyribonucleic acid |
| DSA |
Donor-specific antibodies |
| EAU |
European Association of Urology |
| EMA |
European Medicines Agency |
| EMB |
Endomyocardial biopsy |
| ESC |
European Society of Cardiology |
| ESOT |
European Society of Organ Transplantation |
| FCXM |
Flow crossmatch |
| FDA |
Food and Drug Administration |
| FEV1 |
Forced expiratory volume in one second |
| GEP |
Gene expression profiling |
| HLA |
Human leukocyte antigen |
| INTERCOMEX |
International Collaborative Microarray Study Extension |
| IFN-γ ELISPOT |
Interferon-γ enzyme-linked immunospot |
| ISHLT |
International Society of Heart and Lung Transplantation |
| IVDMIA |
In vitro diagnostic multivariate index assay |
| KDIGO |
Kidney Disease: Improving Global Outcomes |
| LDT |
Laboratory-developed test |
| LFT |
Liver function test |
| MFI |
Mean fluorescence intensity |
| MHC |
Major histocompatibility complex |
| NGS |
Next-generation sequencing |
| NPV |
Negative predictive value |
| OI |
Other injury |
| PBMC |
Peripheral blood mononuclear cells |
| PPV |
Positive predictive value |
| qRT-PCR |
Quantitative real-time polymerase chain reaction methodology |
| RA |
Renal Association |
| RNA |
Ribonucleic acid |
| rsEMB |
Routine surveillance endomyocardial biopsy |
| SAB |
Single-antigen bead |
| STA |
Stable |
| subAR |
Subclinical acute rejection |
| TCMR |
T cell-mediated rejection |
| TLC |
Total lung capacity |
| VOCsVOC |
Volatile organic compound |
Rationale
Solid organ transplant is a delicate process, requiring much oversight and evaluation of every party involved. Rejection, or failure of the transplant, is a potential outcome of any transplant case. At the molecular level, rejection is primarily caused by a component of the adaptive immune system, the major histocompatibility complex (MHC) proteins. These proteins must match between donor and recipient, or the transplant can fail.1
The primary function of MHC proteins is acting as the platform on which T cells identify antigens. Typically, these MHC proteins bind foreign antigens, which are then recognized as such by T cells. From there, the T cells can generate an immune response to handle the antigen. However, the MHC protein products must be identified as “self” by these T cells as well. If an organ donor’s MHC protein does not match the recipient’s, the recipient’s T cells may identify the MHC of the donated organ as “foreign” and subsequently implement an immune response. This eventually starts the cascade of events that causes the transplant to fail.1
Numerous methods mitigate this immune response. Immunosuppressants, which cause desensitization of the immune response, and more have been proposed as methods to circumvent this immune response.1
Proprietary Tests – Gene Expression
AlloMap®
The AlloMap test is based on quantitative real-time polymerase chain reaction methodology (qRT-PCR) using RNA purified from peripheral blood mononuclear cells (PBMC). It is a gene expression test (11 informative genes, nine control genes) that proposes to “aid in the identification of heart transplant recipients with stable allograft function who have a low probability of moderate/severe acute cellular rejection (ACR) at the time of testing in conjunction with standard clinical assessment.”5 Its FDA approval states that it is intended for heart transplant recipients, 15 years or older, and ≥55 days post-transplant.5 CareDX, Inc., the makers of AlloMap states AlloMap can “identify patients with stable allograft function and low probability of cellular rejection.”6
MMDx®
Gene expression profiles available through One Lambda for assessment of transplant rejection include MMDx Heart (1283 genes for heart transplants) and MMDx Kidney (1494 genes for kidney transplants). Whereas other tests are performed on a peripheral blood sample, MMDx Heart and MMDx Kidney requires a biopsy sample. The mRNA levels of a set number of genes (depending on the organ) are assessed in the biopsied tissue and are then compared to the mRNA levels from a reference set of biopsies.7
TruGraf® (Kidney and Liver)
The TruGraf assay is available for kidney and liver transplant recipients. TruGraf gene expression panels are intended for transplant patients and are based on microarray analysis of peripheral blood. TruGraf identifies if a patient is “immune activating” (potentially rejecting) or “immune quiescent” (stable), allowing a clinician to evaluate potential pre-symptomatic kidney damage without use of a biopsy.8,9 According to Eurofins, “120 genes with the highest discriminatory power between the non-TX [“inadequately immunosuppressed”] and TX [“adequately immunosuppressed”] phenotypes are the IP incorporated in TruGraf.”10
Proprietary Tests – Donor-derived Cell-free DNA & Combination
Tissue gene expression is not the only medium tested for rejection. The use of cell-free DNA has shown much promise as a minimally invasive detection method for allograft rejection, and may be used to complement or, ultimately, replace tissue biopsies in the future.11
Viracor TRAC®
The Viracor TRAC® (Transplant Rejection Allograft Check) donor-derived cell-free DNA (dd-cfDNA) assay uses a non-invasive liquid biopsy to monitor the percentage of dd-cfDNA in transplant recipient plasma post-transplant using next generation sequencing (NGS). The test uses “a bioinformatics pipeline that analyzes NGS and genome-wide recipient genotype data to determine the percentage of dd-cfDNA present, which strongly correlates with allograft injury due to rejection. The licensed algorithm accurately quantifies the donor-derived fraction of cell-free DNA without need for donor genotype information, even when the donor is closely related (e.g., sibling kidney donors). The design of this test also enables the determination of dd-cfDNA concentrations over a much wider dynamic range than currently possible with other methods.”12 As of 2023, this technology has not been approved by the FDA. Originally TRAC® was also offered as part of a dual-marker rejection panel by the name of OmniGraf; however, this test is no longer commercially available as of April 1, 2023.
AlloSure® Kidney
AlloSure Kidney, a test offered by the same parent company as AlloMap, evaluates cell-free DNA (cfDNA) in the blood for kidney transplant patients. The test states that when graft injury occurs, donor-derived cell-free DNA is released into the blood where it can be measured as a marker of kidney transplant surveillance. Bloom, et al. (2017) evaluated AlloSure Kidney with 102 kidney recipients. They concluded that a donor-derived cell-free DNA (dd-cfDNA) level of >1% indicated active rejection of the graft.13 According to the manufacturer, AlloSure Kidney is validated for use in stable and high-risk patients, patients with allograft rejection, repeat kidney transplant patients, and pediatric patients.14
A 2021 study from Pai et al. examined the clinical rationale behind a routine testing schedule utilizing dd-cfDNA. In this study, they found that dd-cfDNA surveillance was effective at months one, two, three, four, six, nine, and 12 during the first year following kidney transplantation and quarterly thereafter. They noted that these time points “coincide with major immunologic transition points after transplantation” and that these times points provide clinicians with “molecular information to help inform decision making.”15
AlloSure® Heart
AlloSure Heart detects graft injury via a blood test; the test detects injury and rejection by measuring the amount of dd-cfDNA in the blood. Allosure Heart is covered by Medicare after 55 days when ordered in conjunction with AlloMap and with clinical suspicion of rejection. Heartcare by CareDx is another proprietary combination test that offers AlloMap gene expression profiling combined with next-generation sequencing to quantify dd-cfDNA in cardiac transplant recipients. This combination test includes an AlloMap score, the variability of the AlloMap score, and the percent of dd-cfDNA (using Allosure Heart) in one comprehensive single assessment of heart health.16
AlloSure® Lung
AlloSure® Lung is another donor-derived cell-free DNA (dd-cfDNA) test used for transplant recipients that is now commercially available.17
As evidence of its prowess, AlloSure cites what they describe as “the first and only multi-center study to clinically validate the use of dd-cfDNA in lung transplantation.”17 In a multicenter, retrospective cohort study, Keller, et al. (2022) collected data from lung transplant recipients within three years of transplant and found that “dd-cfDNA was higher in patients with ALAD [acute lung allograft dysfunction] than stable patients” and that the estimated sensitivity of dd-cfDNA ≥1% was 73.9%, specificity of 87.7%, positive predictive value of 43.4% and negative predictive value of 96.5%.” Consequently, the authors concluded that “dd-cfDNA identified acute lung allograft dysfunction in asymptomatic lung transplant patients that may not have been identified by using a clinically indicated biopsy strategy alone,” and as such may “support its use as a potential noninvasive marker for surveillance monitoring.”18
Prospera™
Natera, a genetic testing company, has developed the Prospera test, which uses dd-cfDNA to assess kidney, heart, and lung transplant rejection. This blood test is now covered by Medicare for all kidney transplant recipients.19
Proprietary Tests – Volatile Organic Compounds
Heartsbreath
Another medium used for assessment of rejection is breath. Heartsbreath is an FDA-approved test that purports to predict the probability of grade 3 rejection in heart transplant patients. The
test detects volatile organic compounds.20 The FDA notes that this test does not replace biopsy and is only intended as an adjunct to biopsies. The breath markers are considered to be markers of oxidative stress.21
Clinical Utility and Validity
Pham, et al. (2010) conducted a randomized study comparing gene expression profiling and endomyocardial biopsies for monitoring heart transplant patients. A total of 602 patients who had undergone cardiac transplantation six months to five years previously were included. Both groups were found to have similar rates of primary outcomes, hazard ratios, and two year all-causes of mortality. Patients monitored with gene expression profiling underwent fewer biopsies. The researchers concluded that “Among selected patients who had received a cardiac transplant more than 6 months previously and who were at a low risk for rejection, a strategy of monitoring for rejection that involved gene-expression profiling, as compared with routine biopsies, was not associated with an increased risk of serious adverse outcomes and resulted in the performance of significantly fewer biopsies.”22
Deng, et al. (2014) evaluated the variability of a heart recipient’s gene expression profiling test (AlloMap) scores. Variability was defined as the “the standard deviation of an individual’s cumulative test scores.” A total of 369 patients from the Invasive Monitoring Attenuation by Gene Expression Profiling (IMAGE) study were included, and “gene expression profiling score variability, but not ordinal scores or scores over threshold, was independently associated with future clinical events.” The hazard ratio for a one unit increase in variability was found to be 1.76.23
Kobashigawa, et al. (2015) conducted a single-center randomized controlled trial to evaluate gene expression profiling (GEP) versus endomyocardial biopsy starting at 55 days post-transplant. Sixty heart transplant patients meeting inclusion criteria were randomized beginning at 55 days post-transplant to either GEP or EMB arms. A positive GEP ≥30 between two and six months, or ≥34 after six months, prompted a follow-up biopsy. The primary end point included a composite of death/retransplant, rejection with hemodynamic compromise or graft dysfunction at 18 months post-transplant. The researchers concluded that “GEP starting at 55 days post-transplant seems comparable with EMB for rejection surveillance in selected heart transplant patients and does not result in increased adverse outcomes. GEP also seems useful to guide corticosteroid weaning.”24
Crespo-Leiro, et al. (2015) assessed the “prognostic utility of within-patient variability of GEP scores in predicting future significant clinical events, the negative predictive value (NPV) and the positive predictive value (PPV) of GEP score variability in predicting future significant clinical events.” A total of 737 patients from the Cardiac Allograft Rejection Gene Expression Observational (CARGO) II trial were included. Estimated prevalence of events was found to be 17%, and events occurred at a median of 391 days after the final GEP test. The authors found that “the GEP variability area under the receiver operator characteristics curve for the prediction of a composite event was 0.72. The NPV for GEP score variability of 0.6 was 97% and the PPV for GEP score variability of 1.5 was 35.4%.” The authors concluded that “The GEP score variability may be used in estimating the likelihood of events of death, re-transplantation or graft dysfunction occurring in patients beyond 315 days post-transplant.”25
Furthermore, Crespo-Leiro, et al. (2016) validated the clinical performance of the gene-expression profiling technology in an independent patient population from the CARGO II study. A total of 399 patients were included. The GEP score ranged from 0-39, and the authors identified the optimal cut-off to be 34. At this score (at ≥6 months after transplant), “95.5% (381/399) of GEP tests were true negatives, 4.5% (18/399) were false negatives, 10.2% (6/59) were true positives, and 89.8% (53/59) were false positives.” Based on 938 paired biopsies, the area under the curve for distinguishing ≥3A rejection was found to be 0.70 and 0.69 for 2-6 months and ≥6 months, respectively. The authors concluded, “[T]he choice of threshold score for practical use of GEP testing should consider overall clinical assessment of the patient's baseline risk for rejection.”2
Fujita, et al. (2017) followed up on the CARGO study by investigating the long-term mortality of 46 patients. They found that 23 patients had an increased AlloMap score 6-9 months after heart transplant whereas the remaining 23 patients had a decreased score. After a median follow-up time of 8.1 years, all-cause mortality was significantly elevated in patients with an AlloMap increase compared with patients with a decreased score. The authors concluded, “Dynamic changes of the AlloMap score between 6 and 9 months after HT [heart transplant] were strongly related to all-cause long-term survival after HT. These results suggest that AlloMap potentially displays a useful tool to estimate the patients' risk for long-term mortality.”26
Carey, et al. (2018) analyzed 18 months of follow-up in a national cohort of 27 dual organ recipients (18 heart-kidney, eight heart-liver, one heart-lung) matched to 54 heart-only recipients for gender, age, and time to first GEP (AlloMap) test. They found that “during the first 90 days post-transplant, the mean GEP score for dual organ recipients was 25.2 ± 9.1, vs. 23.5 ± 7.7 for heart-only recipients (P = 0.48), with final GEP scores being 29.1 ± 6.1 and 32.3 ± 3.4, respectively (P = 0.34). GEP scores increased over time at a similar rate (P = 0.33) for both groups. During follow-up, mean GEP score among patients with cytomegalovirus infection was 32.3 (n = 14), compared to 26.7 in patients without cytomegalovirus. Only four (2%) of 233 biopsies were positive for mild antibody-mediated rejection; all occurring in two heart-only recipients (GEP scores = 18-33).”3
Bakir, et al. (2018) analyzed time-dependent phenomapping of clinical and molecular data sets from 94 heart transplant patients (1557 clinical encounters) to determine its accuracy in guiding clinical management. Phenomapping’s associations were analyzed with “immunosuppression therapy, biomarkers, and the combined clinical end point of death, allograft loss, retransplantation, and rejection,” and these findings were further correlated with “clinical parameters, human leucocyte (sic) antigen antibody titers, and peripheral blood mononuclear cell gene expression of the AlloMap test genes.”27 The authors found that patients in the group with higher event rates had “increased human leukocyte antigen class I and II antibody titers, higher expression of the FLT3 AlloMap gene, and lower expression of the MARCH8 and WDR40A AlloMap genes.” The authors concluded that “time-dependent precision phenotyping is a mechanistically insightful, data-driven approach to characterize patterns of clinical care and identify ways to improve clinical management and outcomes.”27
Phillips, et al. (2004) evaluated another novel marker of heart transplant rejection: volatile organic compounds (VOCs). A total of 1061 samples were taken from 539 patients prior to endomyocardial biopsy. The combination of nine VOCs in the algorithm “identified Grade 3 rejection (sensitivity 78.6%, specificity 62.4%, cross-validated sensitivity 59.5%, cross-validated specificity 58.8%, positive predictive value 5.6%, negative predictive value 97.2%). Site pathologists identified the same cases with sensitivity of 42.4%, specificity 97.0%, positive predictive value 45.2% and negative predictive value 96.7%.”28 The authors concluded that “a breath test for markers of oxidative stress was more sensitive and less specific for Grade 3 heart transplant rejection than a biopsy reading by a site pathologist, but the negative predictive values of the 2 tests were similar.”28 However, the Centers for Medicare and Medicaid determined that the evidence does not adequately define the technical characteristics of the test nor demonstrate that Heartsbreath testing to predict heart transplant rejection improves health outcomes.29
Agbor-Enoh, et al. (2019) assessed the donor-derived cell-free DNA (ddcfDNA or dd-cfDNA) levels in 106 lung transplant patients and monitored them for development of allograft failure (“defined as severe chronic lung allograft dysfunction [CLAD], retransplantation, and/or death from respiratory failure”). The average level of donor-derived cell-free DNA (%ddcfDNA) was measured and correlated with allograft failure. The authors separated the patients into three tertiles, with median values of 3.6% in the highest tertile, 1.6% in the middle, and 0.7% in the lowest. The highest tertile was calculated to have a 6.6-fold higher risk of allograft failure compared to the lowest and middle tertiles. The researchers concluded, “lung transplant patients with early unresolving allograft injury measured via %ddcfDNA are at risk of subsequent allograft injury, which is often clinically silent, and progresses to allograft failure.”30
A 14-center post-transplant longitudinal study by Bromberg, et al. (2017) published in The Journal of Applied Laboratory Medicine measured the dd-cfDNA at one, two, three, four, six, nine, and 12 months post-transplant. A total of 380 blood samples were taken during the study, and the median dd-cfDNA value was 0.21%. A value of 1.20% is at the 97.5th percentile. The authors conclude, “In a renal transplant recipient, a dd-cfDNA level above 1.2% is out of range and potentially abnormal.”31
Huang, et al. (2019) evaluated the ability of cell-free DNA to detect rejection in kidney transplant patients. A total of 63 kidney transplant patients with suspicion of rejection were included. Twenty-seven of these had donor-specific antibodies, and 34 were considered to have rejection by biopsy. The percentage of donor-specific cell-free DNA (dd-cfDNA) was higher in patients with antibody-mediated rejection (AMR) compared to those with no rejection and cell-mediated rejection (AMR: 1.35%, no rejection: 0.38%, cell-mediated rejection: 0.27%). A dd-cfDNA percentage of 0.74% was found to yield a sensitivity of 100%, a specificity of 71.8%, a positive predictive value of 68.6%, and a negative predictive value of 100%. The authors concluded that “the dd-cfDNA test did not discriminate CMR from no rejection among kidney transplant recipients, although performance characteristics were stronger for the discrimination of [AMR].”32
Schutz, et al. (2017) evaluated graft-derived cell-free DNA (GcfDNA)’s ability as a marker for liver transplant rejection. A total of 115 patients were included, and 17 patients contributed samples (n = 31) during a biopsy-proven rejection episode; the remaining 88 contributed samples (n = 282) during stable periods. The samples from the rejection cohort were found to have a higher percentage of GcfDNA than the stable cohort (29.3% vs 3.3%). Liver function tests (LFTs) had low correlation rates with GcfDNA, and the area under the curve was 97.1% for GcfDNA. Overall, the authors concluded that “in this study, determination of GcfDNA allowed for earlier and more sensitive discrimination of acute rejection in liver transplant patients as compared with conventional LFTs.”33
A meta-analysis focusing on donor-derived cell-free DNA as a diagnostic tool in transplantation conducted by Oellerich, et al. (2022) reported that the clinical validity of dd-cfDNA in kidney transplantation has been documented in over 20 studies, and that the exponential decline to a baseline level two weeks after transplantation can be used to discriminate graft injury. Moreover, “Although different concepts have been used to establish threshold values for detection of rejection median dd-cfDNA values in reference populations were very similar ranging from 0.21% to 0.40%,” as such consistency is imputed to be evidence of its validity.34 Elevations of dd-cfDNA have also been observed in patients with TCMR and AMBR, and as such subclinical detection may enable earlier therapeutic interventions and improved outcomes.34-37
Grskovic, et al. (2016) performed a validation of AlloSure. The authors included 1117 samples, and AlloSure was used to quantify the fraction of donor-derived cell-free DNA (dd-cfDNA) in both related and unrelated donor-recipient pairs. The quantifiable range was found to be linear from 0.2% to 16%, and the across-runs coefficient of variation was found to be 6.8%. The limit of blank was found to be 0.10%, limit of detection was 0.16%, and limit of quantification was 0.20%. The authors concluded that “application of the assay to clinical samples from heart transplant recipients demonstrated increased levels of dd-cfDNA in patients with biopsy-confirmed rejection and decreased levels of dd-cfDNA after successful rejection treatment.”38
Crespo-Leiro, et al. (2015) compared the levels of dd-cfDNA in heart transplant recipients with biopsy-confirmed rejection to recipients without rejection. A total of 151 plasma samples from 63 patients were evaluated, and 132 of these samples were biopsied. An AlloMap score was also taken. The dd-cfDNA levels were found to be higher in patients with rejection (1.7% vs 0.99%), and an area under curve was measured to be 0.68. The mean AlloMap score was found to be 24.3 in non-rejection patients and 28.3 for rejection patients. The authors found that the dd-cfDNA levels and AlloMap score were not significantly correlated, proposing that these tests may be complementary. Combining the AlloMap and plasma dd-cfDNA levels yielded an AUC of 0.78.39
Jordan, et al. (2018) investigated the use of dd-cfDNA alongside donor-specific antibodies (DSA) testing in identifying antibody-mediated rejection (AMR) of renal allograft recipients (n = 87 patients). They note that the median level of dd-cfDNA was 2.9% in DSA+ patients who have active AMR whereas the dd-cfDNA was significantly lower in both DSA+ patients without AMR (0.34%) or DSA- patients (0.29%). “The positive predictive value of dd-cfDNA (at 1%) to detect active ABMR in DSA+ patients was 81%, whereas the negative predictive value was 83%. The positive predictive value for DSA+ alone was 48%... The combined use of dd-cfDNA and DSA testing may improve the noninvasive diagnosis of active ABMR in kidney transplant patients. Patients with dd-cfDNA+/ DSA+ results have a high probability of active ABMR.”40
Gielis, et al. (2020) obtained samples from 107 kidney transplant recipients to investigate the role of cell-free DNA in acute kidney rejection. Samples were collected between one day and three months after transplantation. The authors noted that increases in cell-free DNA “above a threshold value of 0.88% were significantly associated with the occurrence of episodes of acute rejection (P = 0.017), acute tubular necrosis (P = 0.011) and acute pyelonephritis (P = 0.032).”41 However, the authors also note that “Although increases in plasma ddcfDNA% are associated with graft injury, plasma ddcfDNA does not outperform the diagnostic capacity of the serum creatinine in the diagnosis of acute rejection.”41
Peabody, et al. (2020) researched the clinical utility of the dd-cfDNA Prospera test by Natera to lower the rate of kidney graft loss. Simulated cases of 154 nephrologists were analyzed for this study; some physicians used dd-cfDNA testing and some did not. Results show that at baseline, there were no differences between primary diagnosis, biopsy decisions, or therapeutic management. However, after use of the cc-cfDNA test, “intervention nephrologists were more likely to arrive at the diagnosis of rejection (OR 4.00, 95% CI 1.93-8.30), make a correct decision on biopsy/transplant center referral (OR 11.07, 95% CI 4.87-25.16), and properly adjust therapeutic management (OR 2.37, 95% CI 1.07-5.24).”42
Sigdel, et al. (2018) evaluated an SNP-based assay’s accuracy in identifying allograft rejection or injury. The assay is intended to identify rejection through measurement of donor-derived cell-free DNA (dd-cf DNA). A total of 193 unique renal transplant patients were included, with a total of 300 plasma samples provided. Of the 300 samples, 217 were biopsy-matched, 38 had active rejection (AR), 72 had borderline rejection (BL), 25 had other injury, and 82 were stable (STA). The authors found that median dd-cfDNA was higher in biopsy-proven AR (2.3%) compared to BL (0.6%), OI (0.7%), and STA (0.4%). The assay was found to discriminate active rejection from non-rejection at an area under curve of 0.87, 88.7% sensitivity, and 72.6% specificity (at a cutoff of 1% dd-cfDNA). Of 13 patients with AR findings after six months, 12 tested positive by the assay. The authors concluded that their data supported the “feasibility of using this assay to detect disease prior to renal failure and optimize patient management in the case of allograft injury.”43
Altug, et al. (2019) performed an analytical validation of a “single-nucleotide polymorphism [SNP]-based donor-derived cell-free [cf] DNA assay for detecting rejection in kidney transplant patients.” This test measured 13962 SNPs and was validated using 66 unique samples with 1064 replicates. The authors measured the cf-DNA fraction in related and unrelated (genetically related) donor-recipient pairs. The authors identified a “limit of blank” of 0.11% and a limit of detection and quantitation of 0.15% for unrelated donors. For related donors, a limit of blank of 0.23% and a limit of detection and quantitation of 0.29% was identified. Other metrics such as precision and linearity were found to be identical for both categories. The coefficient of variance was found to be 1.8%. The authors concluded that their findings were an adequate analytical validation of the assay.44
In a retrospective, observational study, Rodgers, et al. (2023) evaluated the correlation, classification, and prediction for AR across two donor-derived cell-free DNA tests and a blood gene-expression profile test in heart transplantation. Looking at patients who had an EMB and both standard and expanded SNP dd-cfDNA testing in the 14 days preceding the EMB, the authors found that “Both standard and expanded SNP dd-cfDNA tests showed no significant difference in testing accuracy for AR,” as the sensitivities of standard SNP and expanded SNP were reported to be 39% and 39%, respectively (p = 0.66). The authors also reported that “Specificity with standard SNP was 82% and expanded SNP testing 84% (p = .26). PPV with standard SNP was 6.2% and expanded SNP testing 7.0% (p = .70). NPV with standard and expanded SNP testing were both 98% (p = .76).” Moreover, the researchers evaluated the screening performance of combining GEP with standard SNP testing (standard dd-cfDNA + GEP), commercially packaged as HeartCare® and determined that “specificity of standard dd-cfDNA + GEP was 47%, which was significantly lower when compared to either standard or expanded SNP dd-cfDNA tests individually (pc < .001).” Regarding the use of GEP in the detection of ACR, “we [the authors] showed GEP specificity was significantly lower than standard SNP dd-cfDNA (pc < .001) but otherwise did not show a significant difference for sensitivity, PPV, and NPV for detection of ACR” and, most notably, “Standard dd-cfDNA + GEP similarly showed significantly reduced specificity versus standard SNP testing alone for detection of ACR (pc < .001).”45 As such, the results suggest that additional coverage offered by expanded SNP assay as compared to the standard SNP assay does “not show a direct impact on discrimination of AR in HTx patients in this study” and “that dd-cfDNA testing, without GEP, is sufficient” for the detection of ACR.45
Henricksen, et al. (2023) conducted a single center, retrospective analysis of adult heart transplant recipients. There, they focused on survival and rejection-free survival one-year post-transplant as primary endpoints for “A GEP cohort, transplanted from January 1, 2015 through December 31, 2017 and eligible for rejection surveillance with GEP was compared to a paired testing cohort, transplanted July 1, 2018 through June 30, 2020, with surveillance from both dd-cfDNA and GEP.” The authors found that “At 1-year, there were no differences between the paired testing and GEP groups in survival (98.4% vs. 94.7%, P = .23) or rejection-free survival (81.3% vs. 73.7% P = .28),” concluding that “Compared to post-transplant rejection surveillance with GEP alone, pairing dd-cfDNA and GEP testing was associated with similar survival and rejection-free survival at 1 year,” with the benefit of requiring significantly fewer biopsies.46
The use of RNA microarray analysis has also begun to garner attention. The INTERCOMEX (International Collaborative Microarray Study Extension) explored the feasibility of real-time MMDx kidney transplant biopsy assessment by comparing the central MMDx assessment with local stand-of-care in 10 experienced North American and European centers that followed Banff 2013 guidelines. A total of 519 samples from 491 patients were deemed sufficient for RNA microarray analysis (3mm), which was used to determine agreement between the histologic diagnoses. The distribution of the principal diagnoses was as follows: antibody-mediated rejection (ABMR, n=88, 17%), ABMR suspected (n=10, 2%), T cell-mediated rejection (TCMR, n=29, 6%), acute kidney injury (AKI, n=43, 8%), “borderline” (n=31, 6%), unexplained atrophy/fibrosis (n=84, 16%), and “no major abnormalities” (n=141, 27%). According to the authors, the report sign-out diagnosis of rejection agreed with histology with a normalized accuracy of 76-77% overall. However, dissecting the accuracy assessment for MMDx yielded PPVs of 51%, 45%, and 90% for ABMR, TCMR, and rejection, respectively, and NPVs of 92%, 96%, and 54% for the same.47 Nevertheless, the authors concluded that “Feedback received for 451 biopsy samples indicates that clinicians agreed with MMDx more often than with their local biopsy assessment and indicated that MMDx would give them more confidence for clinical management.”47
Alam, et al. (2022) evaluated whether MMDx performs in agreement with other validated modalities. Two hundred and twenty-eight corresponding endomyocardial biopsies and MMDx specimens from 135 adult heart transplant patients were retrospectively reviewed with correlating donor-derived cell-free DNA (dd-cfDNA). “Rejection was classified on endomyocardial biopsies in 29 specimens (TCMR ≥ 2R and/or AMR ≥ 1), on MMDx in 56 specimens, and in 74 values with dd-cfDNA ≥0.20%. Despite moderate agreement between endomyocardial biopsies and MMDx (84% agreement, Cohen's kappa, 0.48, p < .001), systematic differences were observed (McNemar's test, p < .001) where MMDx classified 32 of 37 discordant cases as rejection. MMDx and dd-cfDNA demonstrated slight agreement (72% agreement, Cohen's kappa, 0.39, p < .001); however, systematic differences were also apparent where MMDx classified 12 of 50 discordant specimens as rejection when dd-cfDNA was <0.20% (McNemar's test, p < .001).”48 Alam, et al. (2022) findings provide insight on the performance of MMDx relative to other modalities in a heart transplant cohort and have implications on the surveillance and workup of allograft rejection in heart transplantation.
More recently, the TruGraf blood test has emerged as another possible alternative to expensive and invasive biopsies. This proprietary microarray-based test relies on specific gene expression in the peripheral blood as a means of providing information on the adequacy of immunosuppression in transplant candidates with stable renal function, who may manifest subclinical acute rejection (subAR). A study conducted at the Northwestern University Comprehensive Transplant Center and five participating clinical centers for the Genomics for Kidney Transplantation Project reported that in the 125 candidates demonstrating stable renal function, the TruGraf classifier boasted a PPV of 86% (95% CI: 83-89%) and an NPV of 28% (95% CI 13-49%).49 In another study, simultaneous blood tests and clinical assessments were conducted on 192 patients from seven transplant centers to evaluate the predictive power of TruGraf v1. Researchers found that the accuracy of TruGraf—defined here as the agreement between TruGraf result and clinical and/or histologic assessment—was 74% (142/192), with an NPV of 90%.50 Given the data, the researchers assert that the use of the TruGraf classifier will eschew unnecessary surveillance biopsies with high confidence and lead to personalization of management in patient care. However, though these results seem promising, the predictive power of TruGraf in patients with graft dysfunction has yet to be evaluated.
Moayedi, et al. (2019) published results from the Outcomes AlloMap Registry (OAR) study, a multi-center prospective observational study. The OAR study was comprised of the largest group of patients (1,504 individuals) who had undergone gene expression profiling (GEP) for surveillance purposes to-date. Patients were aged ≥ 15 years and ≥ 55 days post-cardiac transplant with a median age of 54 years old. Results indicated the prevalence of “moderate to severe acute cellular rejection (≥2R) was 2.0% from 2 to 6 months and 2.2% after 6 months.” Post-transplant survival at one, two, and five years was 99%, 98%, and 94%, respectively. The authors concluded that patients selected for GEP surveillance had excellent survival outcomes as well as low rates of acute rejection, graft dysfunction, readmission, and death.51
As part of the D-OAR (utility of donor-derived cell free DNA in association with gene expression profiling) trials, Khush, et al. (2019) reported overall that “dd‐cfDNA levels differed significantly between patients with and without AR,” such that the “median level of dd‐cfDNA in patients with AR was significantly higher (0.17%) than in the group of patient specimens without rejection (0.07%, P < .001)” but that “Median dd‐cfDNA levels were 0.17% for both ACR and AMR.”52 When considering only the planned biopsy samples, the authors found that “Statistically different median dd‐cfDNA levels were observed in patients with and without rejection” (p = 0.02, n = 254), while the surveillance biopsy analysis sample size (n = 587) was purportedly “too small to detect a statistically significant difference in median dd‐cfDNA levels” (p =0.140).52 Consequently, the researchers conclude that “We [the authors] demonstrated that dd‐cfDNA levels are significantly higher in patients with AR compared with patients with no biopsy evidence of rejection” and that “Surveillance with dd‐cfDNA may therefore detect early rejection and thereby trigger augmentation of immunosuppression to prevent a more severe rejection event that may result in irreversible graft damage.” However, it should also be noted that many patients were enrolled later than two months posttransplant and that the technique used to evaluate dd‐cfDNA test performance (i.e., endomyocardial biopsy) is not a true “gold standard” for the diagnosis of AR.52
Knüttgen, et al. (2022) analyzed dd-cfDNA percentage in 87 heart transplant recipients. The study goal was to determine the clinical validity of using graft-derived cell-free DNA as well as the association of dd-cfDNA measurements with cardiac allograft rejection. Per the results, sensitivity was 76% and specificity was 83% for cardiac rejection with approximately a 95% confidence interval. Overall, the authors concluded that dd-cfDNA plasma values were “significantly associated with cardiac rejection,” but that pericardial effusions and/or improper sampling “should be considered” as having the ability to confound results in a clinical setting.53
Kamath, et al. (2022) evaluated 72 adult heart transplant patients with at least three concurrent AlloMap/AlloSure results in a single-center retrospective study. The aim of the study was to evaluate the performance of AlloSure, AlloMap, and a combination approach in predicting mortality. During the 480-day follow-up period, five patients died. Higher AlloSure variability (HR 1.66, 95%CI 1.14 – 2.41) was associated with increased mortality risk. However, there was not a strong association (of mortality) with higher AlloMap variability, or of combination AlloSure/AlloMap variability. The authors concluded that, “increased variability of dd-cfDNA in heart transplant patients is associated with both mortality risk and the presence of donor specific antibodies.”54
An observational study by Kim, et al. (2022) aimed to capture the performance of the Prospera dd-cfDNA test in detecting AR in adult heart transplant recipients. The authors reported that “Median dd-cfDNA fraction was significantly higher in samples with a matched biopsy showing AR (median 0.58%, IQR, 0.13%–1.68%) compared to samples where matched biopsies did not show AR (median 0.04%, IQR, 0.01–0.11%, pc < 0.001.” This was confirmed for this study population through ROC analysis, where “the AUC-ROC of the dd-cfDNA test to distinguish AR from non-AR was 0.86” and using a dd-cfDNA fraction threshold of 0.15%, “dd-cfDNA detected histologically-proven rejection with a sensitivity of 78.5% (95% CI, 60.7%–96.3%) and a specificity of 76.9% (95% CI, 71.1%–82.7%). A PPV of 25.1% (95% CI, 18.8%–31.5%) and NPV of 97.3% (95% CI, 95.1%–99.5%) were calculated for this cohort.55 However, the authors acknowledge that “there may have been a selection bias for patients with previous history or suspicion of AMR, who would be more frequently biopsied at our [the chosen] centers,” the higher prevalence of AR in the cohort making it difficult to generalize to patient cohorts with a different risk of rejection, and the limited number of sequential samples.55
Other techniques may be also be worth noting in the future. Montero, et al. (2019) analyzed 12 studies and 1181 patients and found that pretransplant interferon-γ enzyme-linked immunospot (IFN-γ ELISPOT) was “significantly associated with increased AR risk (odds ratio: 3.29; 95% confidence interval (CI), 2.34-4.60); hierarchical summary receiver operating characteristic jointly estimated sensitivity and specificity values were 64.9% (95% CI, 53.7%-74.6%) and 65.8% (95% CI, 57.4%-73.5%), respectively, with moderate heterogeneity across studies.” But though this is the case, “Prospective randomized clinical trials are warranted” given the “suboptimal predictive ability at an individual level.”56
Holzhauser, et al. (2023) reviewed the evidence supporting both GEP and dd-cfDNA testing in heart transplant rejection and provided practical guidance for clinicians to allow for a transition to less invasive options for monitoring rejection after heart transplantation, as well as answering some of the common questions about GEP and dd-cfDNA. One important question they answered was when GEP and dd-cfDNA monitoring should begin post-transplant. As mentioned by previous sources, 55 days post-heart transplant is the earliest timepoint at which GEP monitoring should begin, as the performance of the GEP can be affected by the administration of corticosteroid therapy. In contrast, the dd-cfDNA assay results stabilize approximately 14 days post-heart transplantation and many centers are using dd-cfDNA surveillance as early as three to four weeks post-transplantation.57
Aubert, et al. (2024) studied the use of donor-derived cell-free DNA for kidney allograft rejection testing. The observational study included 2,882 kidney allograft recipients from 14 centers. Overall, dd-cfDNA levels were strongly correlated with allograft rejection, measured by antibody-mediated rejection (P<0.0001), T cell-mediated rejection (P<0.0001), and mixed rejection (P<0.0001). Multivariate analysis also showed that “circulating dd-cfDNA was significantly associated with allograft rejection (odds ratio 2.275; 95% confidence interval (CI) 1.902-2.739; P < 0.0001) independently of standard of care patient monitoring parameters.” Additionally, dd-cfDNA had a high predictive value for the detection of subclinical rejection in stable patients. Overall, the authors concluded that the study “provides insights on the potential value of assessing dd-cfDNA, in addition to standard of care monitoring, to improve the detection of allograft rejection.”58
International Society of Heart and Lung Transplantation
In 2023, the International Society of Heart and Lung Transplantation issued guidelines for the care of heart transplant recipients.59 Below are the recommendations relevant to this policy.
- “It is reasonable to utilize EMB in a HT candidate suspected of having an infiltrative cardiomyopathy or an inflammatory process, such as giant cell myocarditis, amyloidosis, or sarcoidosis. Class IIa, Level of Evidence C
- It is reasonable to perform periodic EMB during the first three to 12 postoperative months for surveillance of rejection Class IIa, Level of Evidence: C
- The standard of care for adolescents should be similar to adults, including surveillance EMB for heart allograft rejection for 3 to 12 months after HT. In younger children, especially infants, the risks associated with EMB and required general anesthesia may outweigh the surveillance benefit for comparably rare acute rejection; therefore, it is reasonable to use a combination of noninvasive screening methods (echocardiography, ECG, biomarkers) instead. Class IIa, Level of Evidence: C
- After the first postoperative year, it is reasonable to continue EMB surveillance in patients who are at higher risk for late acute rejection. This group includes HT recipients with donor-specific antibodies (DSA), a history of recurrent acute rejection, calcineurin-inhibitor free immunosuppression, reduced immunosuppression due to post-transplant malignancy or chronic infection, African American descent. Class IIa, Level of Evidence: C
- Routine EMB later than 5 years after HT are not recommended. EMB should be performed only for cause in patients with signs or symptoms of cardiac allograft dysfunction. Class III, Level of Evidence: C
- Children receiving ABO incompatible cardiac allografts in the first 2 years of life with isohemagglutinin titers toward the donor blood group below 1:32 and without elevated titers post-transplant do not require more frequent EMB or non-invasive monitoring compared to recipients of ABO compatible organs. Class IIa, Level of Evidence: B
- Ventricular evoked responses (VER) monitoring for rejection surveillance is no longer recommended as the technology has become obsolete. Class III, Level of Evidence: C
- Gene Expression Profiling (GEP) (i.e., Allomap) of peripheral blood can be used in low-risk patients between 2 months and 5 years after HT to identify adult recipients who have low risk of current ACR to reduce the frequency of EMB. Data in children does not allow a general recommendation of GEP as a routine tool at present. Class IIa, Level of Evidence: B
- It is reasonable to integrate biomarkers such as BNP and high-sensitivity troponins into a rejection monitoring strategy to identify higher risk patients who may benefit from additional evaluation for ACR, AMR, or CAV.
- Class IIb, Level of Evidence C
- Post-transplant monitoring for DSA should be performed at 1, 3, 6, and 12 months postoperatively and annually thereafter. Sensitized patients should be monitored more frequently. Class IIa, Level of Evidence C
- The use of systemic inflammatory markers such as C-reactive protein (CRP) for acute heart allograft rejection monitoring is not recommended. Class III, Level of Evidence: C
- In younger children, especially infants, the risks associated with EMB and required general anesthesia may outweigh the surveillance benefit for comparably rare acute rejection; therefore, it is reasonable to use a combination of noninvasive screening methods (echocardiography, ECG, biomarkers) instead. Class IIa, Level of Evidence: C
- Use of the immune cell function assay (ImmuKnow) cannot be recommended in adult and pediatric heart transplant recipients for rejection monitoring. Class III, Level of Evidence B
- Patient Management During a Pandemic: Efforts should be made to reduce visits by clinically stable heart transplant patients to medical facilities by shifting blood testing to the patients’ homes. Remote drawing of blood samples can include screening tests to determine if patients require endomyocardial biopsies using gene expression profiling and donor derived cell-free DNA assays. Class I, Level of Evidence C.”59
In a 2016 guideline discussing antibody-mediated rejection (AMR) of the lung, the ISHLT noted the lack of specific diagnostic criteria for AMR and listed allograft dysfunction, positive histology, positive C4d staining, and donor-specific anti–human leukocyte antigen (HLA) antibodies (DSA) as potential diagnostic items for AMR.60
Regarding chronic lung allograft dysfunction (CLAD), the society recommended the following:
- “In stable CLAD patients with stable FEV1 [forced expiratory volume in 1 second] or a very slow decline in FEV1 (so-called plateau phase), it is advised to have lung function measured at least every 3 or 4 months.”
- “We recommend measuring TLC [total lung capacity] in LTx patients at 3 and 6 months after transplant and annually thereafter. TLC measurements should also be obtained if changes ≥10% from previous values. The “gold standard” technique to assess TLC is body plethysmography.”
- “The initial CT scans (inspiratory views with a maximum width of 3-mm sections, and expiratory sections as well) without contrast media are recommended in all LTx patients at 6-month follow-up (when spirometry is usually optimal). Repeat CT studies should be obtained when CLAD is initially diagnosed to better visualize air trapping and various subtle opacities.”
- “Transbronchial biopsy and bronchoalveolar lavage (BAL) have a major role in the detection of treatable causes prior to the diagnosis of definite CLAD and should be performed at the start of the diagnostic process to investigate the decline in lung function not explained by obvious, non-CLAD causes.”
- “BAL should also be assessed for signs of aspiration, which is suggested by the presence of multinucleated giant cells or foreign organic material (such as meat and plant material), and/or of lipid, as demonstrated on a lipid stain such as oil red-O or Sudan black and/or bile acids (detected by enzymatic assay).”61
In 2024, the ISHLT published a guideline on the evaluation and care of cardiac transplant candidates, stating: “HLA antibody screening is performed for every patient before listing and while listed for transplantation every 3 to 6 months by most centers following the technical and interpretation guidance of the STAR working group recommendations. Donor hearts are not selected on the basis of HLAs because of time restrictions related to cardiac preservation; therefore, tissue type should be determined for retrospective analysis and may assist with determination of donor-specific antibodies. Flow cytometry is an immunofluorescence method for identifying cell surface antigens by detecting conjugated antibody. Results from flow cytometry allow for assessment regarding the risk of a positive crossmatch at the time of transplant. Patients at risk for suboptimal outcome post-transplant are defined as having a PRA > 10% or donor-specific antibodies at the time of transplantation. Decisions can be made with more confidence regarding the need for a prospective vs retrospective crossmatch, as well as giving providers more insight into the likelihood of antibody-mediated rejection after transplantation.”62
American Society of Transplant Surgeons
A positional statement on the role of dd-cfDNA in transplant surveillance was approved by the ASTS Executive Committee in March of 2023. The statement aims to provide clarity for clinicians and to advocate for access to molecular diagnostic techniques to support improvement in long term allograft function. ASTS recommends the following, particularly for adult individuals:
- “We suggest that clinicians consider measuring serial dd-cfDNA levels in kidney transplant recipients with stable renal allograft function to exclude the presence of subclinical antibody mediated rejection.
- We recommend that clinicians measure dd-cfDNA levels in kidney transplant recipients with acute allograft dysfunction to exclude the presence of rejection, particularly antibody-mediated rejection (ABMR).
- We do not recommend the use of blood gene expression profiling (GEP) in kidney transplant recipients for the purpose of diagnosing or excluding sub-clinical rejection, as adequate evidence supporting such use is still lacking.
- We do not recommend the use of blood GEP to diagnose or exclude the presence of acute graft rejection in kidney transplant recipients with acute allograft dysfunction given the paucity of data to support this practice.
- We recommend that dd-cfDNA may be utilized to rule out subclinical rejection in heart transplant recipients.
- We recommend that clinicians utilize peripheral blood GEP as a non-invasive diagnostic tool to rule out acute cellular rejection in stable, low-risk, adult heart transplant recipients who are over 55 days status post heart transplantation.
- We recommend that there is still insufficient evidence to recommend dd-cfDNA or GEP testing in liver transplant recipients.”63
American Society of Transplantation
The AST’s Thoracic and Critical Care Community published a review of currently available and emerging biomarkers to aid in the management of heart transplant recipients. The panel expressed a need for larger and more sophisticated studies to rigorously validate the growing number of potentially useful biomarkers in heart transplant medicine; this need is underscored by the growing awareness that the current gold-standard for detecting rejection (endomyocardial biopsy) has several limitations.
Based on the currently available data, over half the members of the expert panel agreed that Donor-specific HLA antibodies (DSA) and dd-cfDNA used in combination may be the optimal biomarkers to assess clinical outcomes for heart transplant recipients. Despite this potential, however, the committee cautions that current dd-cfDNA assays lack tissue specificity, cannot distinguish between ACR and AMR, and have unproven clinical utility in the context of heart transplant medicine (due to a lack of clinical utility trials).64
Heart Failure Association of the European Society of Cardiology (ESC)
The Heart Failure Association of the European Society of Cardiology published a position statement on Advanced Heart Failure which states: “Post-transplant patients should undergo a pre-defined regimen of graft biopsies, titration of immunosuppressive and other therapies, rejection monitoring, assessment for infections, transplant coronary artery disease and/or cardiac allograft vasculopathy, immunosuppression side effects, and other potential complications including neoplasia, and co-morbidities that require comprehensive treatment.”65
A joint position statement on endomyocardial biopsy by the Heart Failure Association, Heart Failure Society of America, and Japanese Heart Failure Society Position elaborates, stating that “EMB remains the gold standard for the detection of HTx rejection. EMB after HTx can be scheduled according to a protocol for routine surveillance EMB (rsEMB) in asymptomatic patients, and it is also performed in patients with worsening clinical status, as a symptom triggered EMB.” However, it is admitted that though this is the case, “there is a lack of consensus on the optimal timing and frequency of rsEMB.”66
European Association of Urology
The European Association of Urology published guidelines on renal transplantation. In it, they state that “the ultimate standard for the diagnosis of rejection is transplant biopsy, because it is impossible to differentiate acute rejection solely on clinical indicators from other causes of renal dysfunction (e.g. acute tubular necrosis, infection, disease recurrence or CNI nephrotoxicity). Therefore, all rejections should be verified by renal biopsy.”67
The EAU also asserted that when it comes to matching donors with recipients, the following recommendations apply:
- “Determine the ABO blood group and the human leukocyte antigen A, B, C and DR phenotypes for all candidates awaiting kidney transplantation.”
- “Test both the donor and recipient for human leukocyte antigen DQ. Human leukocyte antigen DP testing may be performed for sensitised patients.”
- “Perform thorough testing for HLA antibodies before transplantation.”
- “Perform adequate cross-match tests to avoid hyper-acute rejection, before each kidney and combined kidney/pancreas transplantation.”67
Renal Association (RA)
The Renal Association published guidelines regarding post-operative care for kidney transplant patients. These guidelines have been endorsed by the British Transplant Society (BTS). The assessment of rejection recommendations is provided below:
- “We recommend that a transplant renal biopsy should be carried out before treating an acute rejection episode unless this will substantially delay treatment or pose a significant risk to the patient.”
- “We recommend that a protocol transplant renal biopsy, defined as a biopsy performed in a stable graft without clinical evidence of acute rejection, be considered in the setting of persisting delayed graft function.”
Furthermore, in the rationale, the RA states that “Rejection episodes are characteristically associated with loss of graft function but diagnosis is best established by a percutaneous biopsy since it differentiates rejection clearly from other causes of graft dysfunction. Recognition of different forms of rejection may inform different treatment regimens (e.g. antibody mediated rejection).”68
American Association for the Study of Liver Diseases (AASLD) and the American Society of Transplantation
These joint guidelines provide guidance on the long-term management of liver transplants. Their recommendations concerning assessment of rejection are as follows:
- “Rejection can be reliably diagnosed only on the basis of liver histology; a biopsy sample should be taken before treatment initiation and classified according to the Banff criteria.”
- The guidelines also note that “Both forms of rejection are, until the late stages, asymptomatic, and the diagnosis is made through the investigation of abnormal liver tests; the diagnosis can be confirmed only on the basis of histology.”69
Kidney Disease: Improving Global Outcomes (KDIGO)
The KDIGO does not list any gene expression or cell-free DNA techniques in their guideline for evaluating and managing transplant recipient patients.70,71
European Society for Organ Transplantation (ESOT)
A working group from the European Society for Organ Transplantation submitted a Broad Scientific Advice request to the European Medicines Agency (EMA) outlining the alloimmune and non-alloimmune factors driving allograft rejection. ESOT proposes the following for alloimmune risk stratification in CDC-negative kidney transplantation:
- Transplantation performed with an HLA-identical donor carries a significantly lower immunological risk than transplantation from a donor of another HLA status.
- For the same allogeneic eplet load, grafts from living donors, which are better preserved and are therefore less immunogenic than grafts from deceased donors, are associated with a lower immunological risk.
- Patients with anti-donor serological memory at the time of or short time before transplantation (i.e., those with the presence of HLA-DSA) should be clearly differentiated from the others:
- Patients with donor reactivity are likely to have immune reactions to the allograft, with a heightened risk of post-transplant AMR and poor allograft outcome.
- SAB testing is the gold standard to establish the repertoire of serologic memory and define the presence of a recipient’s circulating anti-HLA DSA.
- Local transplant physicians and immunologists should determine the appropriate cut-off point, with a focus on plausibility of immunization history.
- In absence of clinical validation, non–HLA-DSA routine screening assays should not be considered when evaluating immunological risk of a transplantation.
- Using SAB testing, three risk groups can be identified (patients with non–donor-specific HLA antibodies and no previous exposure to donor antigen are considered as naïve patients):
- Group 1: Patients with no signs of anti-HLA immune sensitization at any time point (very low risk).
- Group 2: Patients with previous exposure to donor antigens or history of HLA-DSA positivity, but without HLA-DSA at time of transplantation (intermediate risk due to likely presence of memory T and/or B alloimmune response): T/B ELISpot assays could identify anti-donor memory cells, but without clinical validation these assays should not be considered when evaluating immunological risk of a transplantation.
- Group 3: Patients with HLA-DSA at time of transplantation (high risk). There is ongoing discussion on the utility of MFI in SAB or FCXM as a potential surrogate to estimate the DSA titer and for individual risk stratification.
- Molecular HLA mismatch analysis is likely to play a future role in better allocating more compatible allografts, as well as in stratifying the primary alloimmune risk. However, in the absence of consensus regarding what algorithm (and which cut-off) should be used to quantify the eplet load and whether the quality of eplet should also be considered, it is difficult to integrate such information immediately into clinical practice and clinical trial design. Further consensus building is necessary.
- Although it is a fast-evolving field, no reliable test is currently available to measure innate immune alloreactivity in routine clinical practice.72
In reviewing these conclusions, the Committee for Medicinal Products for Human Use (CHMP) of the European Medicines Agency (EMA) “agreed that several important issues need to be considered in assessing the alloimmune risk following kidney transplantation. These include general characteristics of the recipient and donor, as well as issues related to the transplanted organ and issues requiring further studies and/or consensus before adapting into general guidelines.”72
References:
- Vella J. Transplantation immunobiology. Updated April 9, 2024. https://www.uptodate.com/contents/transplantation-immunobiology
- Crespo-Leiro MG, Stypmann J, Schulz U, et al. Clinical usefulness of gene-expression profile to rule out acute rejection after heart transplantation: CARGO II. European heart journal. Sep 01 2016;37(33):2591-601. doi:10.1093/eurheartj/ehv682
- Carey SA, Tecson KM, Jamil AK, Felius J, Wolf-Doty TK, Hall SA. Gene expression profiling scores in dual organ transplant patients are similar to those in heart-only recipients. Transplant immunology. Mar 26 2018;doi:10.1016/j.trim.2018.03.003
- Gielis EM, Ledeganck KJ, De Winter BY, et al. Cell-Free DNA: An Upcoming Biomarker in Transplantation. Am J Transplant. Oct 2015;15(10):2541-51. doi:10.1111/ajt.13387
- FDA. 510(k) Substantial Equivalence Determination. https://www.accessdata.fda.gov/cdrh_docs/reviews/k073482.pdf
- CareDX. Identify Heart Transplant Injury With HeartCare. https://caredx.com/products-and-services/transplant-services/heart/heartcare/
- One Lambda. MMDx® for Molecular Biopsy Assessment. https://www.thermofisher.com/onelambda/us/en/post-transplant/molecular-biopsy-assessment.html
- Eurofins Viracor. TruGraf® Blood Gene Expression Test Test Code: 30929. https://www.eurofins-viracor.com/
- Peddi VR, Patel PS, Schieve C, Rose S, First MR. Serial Peripheral Blood Gene Expression Profiling to Assess Immune Quiescence in Kidney Transplant Recipients with Stable Renal Function. Ann Transplant. Apr 28 2020;25:e920839. doi:10.12659/aot.920839
- Eurofins. How Do You Anticipate Allograft Rejection? https://transplantgenomics.com/biomarkers/
- Pattar S, Greenway S. Monitoring the Health of Solid Organs After Transplantation Using Cell-Free DNA. AACC. 2020;
- eurofins. Viracor TRAC® Kidney dd-cfDNA. https://www.eurofins-viracor.com/
- Bloom RD, Bromberg JS, Poggio ED, et al. Cell-Free DNA and Active Rejection in Kidney Allografts. Journal of the American Society of Nephrology : JASN. Jul 2017;28(7):2221-2232. doi:10.1681/asn.2016091034
- CareDX. AlloSure Kidney Care Test Info. https://caredx.com/products-and-services/transplant-services/kidney/allosure/
- Pai A, Swan JT, Wojciechowski D, et al. Clinical Rationale for a Routine Testing Schedule Using Donor-Derived Cell-Free DNA After Kidney Transplantation. Ann Transplant. Jul 2 2021;26:e932249. doi:10.12659/AOT.932249
- CareDx. HeartCare https://caredx.com/products-and-services/transplant-services/heart/heartcare/
- CareDx. Allosure Lung. https://caredx.com/products-and-services/transplant-services/lung/
- Keller M, Sun J, Mutebi C, et al. Donor-derived cell-free DNA as a composite marker of acute lung allograft dysfunction in clinical care. J Heart Lung Transplant. Apr 2022;41(4):458-466. doi:10.1016/j.healun.2021.12.009
- Natera. Prospera. https://www.natera.com/organ-health/prospera-organ-transplantation-assessment
- FDA. The Heartsbreath Test: Product Labeling for Physicians. https://www.accessdata.fda.gov/cdrh_docs/pdf3/H030004c.pdf
- FDA. Summary of Safety and Probable Benefit. . U.S. Food and Drug Administration. https://www.accessdata.fda.gov/cdrh_docs/pdf3/H030004B.pdf
- Pham MX, Teuteberg JJ, Kfoury AG, et al. Gene-expression profiling for rejection surveillance after cardiac transplantation. The New England journal of medicine. May 20 2010;362(20):1890-900. doi:10.1056/NEJMoa0912965
- Deng MC, Elashoff B, Pham MX, et al. Utility of gene expression profiling score variability to predict clinical events in heart transplant recipients. Transplantation. Mar 27 2014;97(6):708-14. doi:10.1097/01.TP.0000443897.29951.cf
- Kobashigawa J, Patel J, Azarbal B, et al. Randomized pilot trial of gene expression profiling versus heart biopsy in the first year after heart transplant: early invasive monitoring attenuation through gene expression trial. Circulation Heart failure. May 2015;8(3):557-64. doi:10.1161/circheartfailure.114.001658
- Crespo-Leiro MG, Stypmann J, Schulz U, et al. Performance of gene-expression profiling test score variability to predict future clinical events in heart transplant recipients. BMC cardiovascular disorders. Oct 09 2015;15:120. doi:10.1186/s12872-015-0106-1
- Fujita B, Prashovikj E, Schulz U, et al. Predictive value of gene expression profiling for long-term survival after heart transplantation. Transplant immunology. Mar 2017;41:27-31. doi:10.1016/j.trim.2017.02.001
- Bakir M, Jackson NJ, Han SX, et al. Clinical phenomapping and outcomes after heart transplantation. J Heart Lung Transplant. Aug 2018;37(8):956-966. doi:10.1016/j.healun.2018.03.006
- Phillips M, Boehmer JP, Cataneo RN, et al. Heart allograft rejection: detection with breath alkanes in low levels (the HARDBALL study). J Heart Lung Transplant. Jun 2004;23(6):701-8.
- CMS. National Coverage Determination (NCD) for Heartsbreath Test for Heart Transplant Rejection (260.10). https://www.cms.gov/medicare-coverage-database/view/ncd.aspx?NCDId=325
- 30. Agbor-Enoh S, Wang Y, Tunc I, et al. Donor-derived cell-free DNA predicts allograft failure and mortality after lung transplantation. EBioMedicine. Feb 2019;40:541-553. doi:10.1016/j.ebiom.2018.12.029
- 31. Bromberg JS, Brennan DC, Poggio E, et al. Biological Variation of Donor-Derived Cell-Free DNA in Renal Transplant Recipients: Clinical Implications. The Journal of Applied Laboratory Medicine: An AACC Publication. 2017;2(3):309-321. doi:10.1373/jalm.2016.022731
- 32. Huang E, Sethi S, Peng A, et al. Early clinical experience using donor-derived cell-free DNA to detect rejection in kidney transplant recipients. Am J Transplant. Jun 2019;19(6):1663-1670. doi:10.1111/ajt.15289
- 33. Schutz E, Fischer A, Beck J, et al. Graft-derived cell-free DNA, a noninvasive early rejection and graft damage marker in liver transplantation: A prospective, observational, multicenter cohort study. PLoS medicine. Apr 2017;14(4):e1002286. doi:10.1371/journal.pmed.1002286
- Oellerich M, Budde K, Osmanodja B, et al. Donor-derived cell-free DNA as a diagnostic tool in transplantation. Front Genet. 2022;13:1031894. doi:10.3389/fgene.2022.1031894
- Oellerich M, Sherwood K, Keown P, et al. Liquid biopsies: donor-derived cell-free DNA for the detection of kidney allograft injury. Nat Rev Nephrol. Sep 2021;17(9):591-603. doi:10.1038/s41581-021-00428-0
- Halloran PF, Reeve J, Madill-Thomsen KS, Demko Z, Prewett A, Billings P. The Trifecta Study: Comparing Plasma Levels of Donor-derived Cell-Free DNA with the Molecular Phenotype of Kidney Transplant Biopsies. Journal of the American Society of Nephrology : JASN. Feb 2022;33(2):387-400. doi:10.1681/asn.2021091191
- Knight SR, Thorne A, Lo Faro ML. Donor-specific Cell-free DNA as a Biomarker in Solid Organ Transplantation. A Systematic Review. Transplantation. Feb 2019;103(2):273-283. doi:10.1097/tp.0000000000002482
- Grskovic M, Hiller DJ, Eubank LA, et al. Validation of a Clinical-Grade Assay to Measure Donor-Derived Cell-Free DNA in Solid Organ Transplant Recipients. The Journal of molecular diagnostics : JMD. Nov 2016;18(6):890-902. doi:10.1016/j.jmoldx.2016.07.003
- Crespo-Leiro M, Zuckermann A, Stypmann J, et al. Increased Plasma Levels of Donor-Derived Cell-Free DNA Correlate With Rejection in Heart Transplant Recipients: The CARGO II Multicenter Trial. The Journal of Heart and Lung Transplantation. 2015;34(4):S31-S32. doi:10.1016/j.healun.2015.01.075
- Jordan SC, Bunnapradist S, Bromberg JS, et al. Donor-derived Cell-free DNA Identifies Antibody-mediated Rejection in Donor Specific Antibody Positive Kidney Transplant Recipients. Transplant Direct. Sep 2018;4(9):e379. doi:10.1097/txd.0000000000000821
- Gielis EM, Ledeganck KJ, Dendooven A, et al. The use of plasma donor-derived, cell-free DNA to monitor acute rejection after kidney transplantation. Nephrol Dial Transplant. Apr 1 2020;35(4):714-721. doi:10.1093/ndt/gfz091
- Peabody J, Billings P, Valdenor C, et al. Randomized clinical trial of a novel donor-derived cfDNA test to detect rejection in CPV-simulated renal transplant patients. Int Urol Nephrol. Aug 2020;52(8):1593-1601. doi:10.1007/s11255-020-02491-1
- Sigdel TK, Archila FA, Constantin T, et al. Optimizing Detection of Kidney Transplant Injury by Assessment of Donor-Derived Cell-Free DNA via Massively Multiplex PCR. J Clin Med. Dec 23 2018;8(1)doi:10.3390/jcm8010019
- Altug Y, Liang N, Ram R, et al. Analytical Validation of a Single-nucleotide Polymorphism-based Donor-derived Cell-free DNA Assay for Detecting Rejection in Kidney Transplant Patients. Transplantation. 2019;103(12)
- Rodgers N, Gerding B, Cusi V, et al. Comparison of two donor-derived cell-free DNA tests and a blood gene-expression profile test in heart transplantation. Clin Transplant. Jun 2023;37(6):e14984. doi:10.1111/ctr.14984
- Henricksen EJ, Moayedi Y, Purewal S, et al. Combining donor derived cell free DNA and gene expression profiling for non-invasive surveillance after heart transplantation. Clin Transplant. Mar 2023;37(3):e14699. doi:10.1111/ctr.14699
- Halloran PF, Reeve J, Akalin E, et al. Real Time Central Assessment of Kidney Transplant Indication Biopsies by Microarrays: The INTERCOMEX Study. Am J Transplant. Nov 2017;17(11):2851-2862. doi:10.1111/ajt.14329
- Alam A, Van Zyl J, Paul Milligan G, Michelle McKean S, Patel R, Anne Hall S. Evolving the surveillance and workup of heart transplant rejection: A real-world analysis of the Molecular Microscope Diagnostic System. Am J Transplant. Oct 2022;22(10):2443-2450. doi:10.1111/ajt.17087
- First MR, Whisenant T, Friedewald JJ, et al. Clinical Utility of Peripheral Blood Gene Expression Profiling of Kidney Transplant Recipients to Assess the Need for Surveillance Biopsies in Subjects with Stable Renal Function. Journal of Transplantation Technologies & Research. 2017;7(3)doi:10.4172/2161-0991.1000177
- Marsh CL, Kurian SM, Rice JC, et al. Application of TruGraf v1: A Novel Molecular Biomarker for Managing Kidney Transplant Recipients With Stable Renal Function. Transplantation Proceedings. 2019/04/01/ 2019;51(3):722-728. doi:10.1016/j.transproceed.2019.01.054
- Moayedi Y, Foroutan F, Miller RJH, et al. Risk evaluation using gene expression screening to monitor for acute cellular rejection in heart transplant recipients. J Heart Lung Transplant. Jan 2019;38(1):51-58. doi:10.1016/j.healun.2018.09.004
- Khush KK, Patel J, Pinney S, et al. Noninvasive detection of graft injury after heart transplant using donor-derived cell-free DNA: A prospective multicenter study. Am J Transplant. Oct 2019;19(10):2889-2899. doi:10.1111/ajt.15339
- Knüttgen F, Beck J, Dittrich M, et al. Graft-derived Cell-free DNA as a Noninvasive Biomarker of Cardiac Allograft Rejection: A Cohort Study on Clinical Validity and Confounding Factors. Transplantation. Mar 1 2022;106(3):615-622. doi:10.1097/tp.0000000000003725
- Kamath M, Shekhtman G, Grogan T, et al. Variability in Donor-Derived Cell-Free DNA Scores to Predict Mortality in Heart Transplant Recipients - A Proof-of-Concept Study. Front Immunol. 2022;13:825108. doi:10.3389/fimmu.2022.825108
- Kim PJ, Olymbios M, Siu A, et al. A novel donor-derived cell-free DNA assay for the detection of acute rejection in heart transplantation. J Heart Lung Transplant. Jul 2022;41(7):919-927. doi:10.1016/j.healun.2022.04.002
- Montero N, Farouk S, Gandolfini I, et al. Pretransplant Donor-specific IFNγ ELISPOT as a Predictor of Graft Rejection: A Diagnostic Test Accuracy Meta-analysis. Transplant Direct. May 2019;5(5):e451. doi:10.1097/txd.0000000000000886
- Holzhauser L, DeFilippis EM, Nikolova A, et al. The End of Endomyocardial Biopsy?: A Practical Guide for Noninvasive Heart Transplant Rejection Surveillance. JACC Heart Fail. Mar 2023;11(3):263-276. doi:10.1016/j.jchf.2022.11.002
- Aubert O, Ursule-Dufait C, Brousse R, et al. Cell-free DNA for the detection of kidney allograft rejection. Nat Med. Aug 2024;30(8):2320-2327. doi:10.1038/s41591-024-03087-3
- Velleca A, Shullo MA, Dhital K, et al. The International Society for Heart and Lung Transplantation (ISHLT) guidelines for the care of heart transplant recipients. J Heart Lung Transplant. May 2023;42(5):e1-e141. doi:10.1016/j.healun.2022.10.015
- Levine DJ, Glanville AR, Aboyoun C, et al. Antibody-mediated rejection of the lung: A consensus report of the International Society for Heart and Lung Transplantation. J Heart Lung Transplant. Apr 2016;35(4):397-406. doi:10.1016/j.healun.2016.01.1223
- Verleden GM, Glanville AR, Lease ED, et al. Chronic lung allograft dysfunction: Definition, diagnostic criteria, and approaches to treatment-A consensus report from the Pulmonary Council of the ISHLT. J Heart Lung Transplant. May 2019;38(5):493-503. doi:10.1016/j.healun.2019.03.009
- Peled Y, Ducharme A, Kittleson M, et al. International Society for Heart and Lung Transplantation Guidelines for the Evaluation and Care of Cardiac Transplant Candidates-2024. J Heart Lung Transplant. Oct 2024;43(10):1529-1628.e54. doi:10.1016/j.healun.2024.05.010
- ASTS. ASTS Statement on donor-derived cell-free DNA (dd-cf-DNA). https://asts.org/docs/default-source/position-statements/dd-cfdna-position-statement.pdf
- Kobashigawa J, Hall S, Shah P, et al. The evolving use of biomarkers in heart transplantation: Consensus of an expert panel. Am J Transplant. Jun 2023;23(6):727-735. doi:10.1016/j.ajt.2023.02.025
- Crespo-Leiro MG, Metra M, Lund LH, et al. Advanced heart failure: a position statement of the Heart Failure Association of the European Society of Cardiology. European journal of heart failure. May 27 2018;doi:10.1002/ejhf.1236
- Seferović PM, Tsutsui H, McNamara DM, et al. Heart Failure Association, Heart Failure Society of America, and Japanese Heart Failure Society Position Statement on Endomyocardial Biopsy. Journal of Cardiac Failure. 2021/07/01/ 2021;27(7):727-743.
- Kranz J, Bartoletti R, Bruyère F, et al. European Association of Urology Guidelines on Urological Infections: Summary of the 2024 Guidelines. Eur Urol. Jul 2024;86(1):27-41. doi:10.1016/j.eururo.2024.03.035
- Baker RJ, Mark PB, Patel RK, Stevens KK, Palmer N. Renal association clinical practice guideline in post-operative care in the kidney transplant recipient. BMC Nephrol. Jun 2 2022;18(1):174. doi:10.1186/s12882-017-0553-2
- Lucey MR, Terrault N, Ojo L, et al. Long-Term Management of the Successful Adult Liver Transplant: 2012 Practice Guideline by AASLD and the AST. 2013;
- KDIGO. Managing KIDNEY TRANSPLANT RECIPIENTS. 2010;
- KDIGO. KDIGO Clinical Practice Guideline on the Evaluation and Management of Candidates for Kidney Transplantation. Official Journal of the Tranplantation Society and International Liver Transplantation Society. 2020;104(4S)
- Bestard O, Thaunat O, Bellini MI, et al. Alloimmune Risk Stratification for Kidney Transplant Rejection. Transpl Int. 2022;35:10138. doi:10.3389/ti.2022.10138
Coding Section
| Codes | Number | Description |
| CPT | 81479 | Unlisted molecular pathology procedure |
| 81558 (effective 01/01/2025) | Transplantation medicine (allograft rejection, kidney), mRNA, gene expression profiling by quantitative polymerase chain reaction (qPCR) of 139 genes, utilizing whole blood, algorithm reported as a binary categorization as transplant excellence, which indicates immune quiescence, or not transplant excellence, indicating subclinical rejection | |
| 81595 (effective 1/1/2016) | Cardiology (heart transplant), mRNA, gene expression profiling by real-time quantitative PCR of 20 genes (11 content and 9 housekeeping), utilizing subfraction of peripheral blood, algorithm reported as a rejection risk score. Proprietary test: AlloMap® Lab/Manufacturer: CareDx Inc. |
|
| 81599 | Unlisted multianalyte assay with algorithmic analysic | |
| 84999 | Unlisted chemistry procedure | |
| 0055U | Cardiology (heart transplant), cell-free DNA, PCR assay of 96 DNA target sequences (94 single nucleotide polymorphism targets and two control targets), plasma Proprietary test: myTAIHEART Lab/Manufacturer: TAI Diagnostics Inc. |
|
| 0087U | Cardiology (heart transplant), mRNA gene expression profiling by microarray of 1283 genes, transplant biopsy tissue, allograft rejection and injury algorithm reported as a probability score |
|
| 0088U | Transplantation medicine (kidney allograft rejection), microarray gene expression profiling of 1494 genes, utilizing transplant biopsy tissue, algorithm reported as a probability score for rejection |
|
| 0118U (effective 10/01/2024) | Transplantation medicine, quantification of donor-derived cell-free DNA using whole genome next-generation sequencing, plasma, reported as percentage of donor-derived cell-free DNA in the total cell-free DNA. | |
| 0319U | Nephrology (renal transplant), RNA expression by select transcriptome sequencing, using pretransplant peripheral blood, algorithm reported as a risk score for early acute rejection Proprietary test: Clarava™ Lab/Manufacturer: Verici Dx/Verici Dx Inc. |
|
| 0320U | Nephrology (renal transplant), RNA expression by select transcriptome sequencing, using posttransplant peripheral blood, algorithm reported as a risk score for acute cellular rejection Proprietary test: Tuteva™ Lab/Manufacturer: Verici Dx/Verici Dx Inc. |
|
| 0493U (effective 10/01/2024) | Transplantation medicine, quantification of donor-derived cell-free DNA (cfDNA) using nextgeneration sequencing, plasma, reported as percentage of donorderived cell-free DNA | |
| 0508U (effective 10/01/2024) | Transplantation medicine, quantification of donor-derived cell-free DNA using 40 single-nucleotide polymorphisms (SNPs), plasma, and urine, initial evaluation reported as percentage of donor-derived cell-free DNA with risk for active rejection | |
| 0509U (effective 10/01/2024) | Transplantation medicine, quantification of donor-derived cell-free DNA using up to 12 single-nucleotide polymorphisms (SNPs) previously identified, plasma, reported as percentage of donor-derived cell-free DNA with risk for active rejection | |
| 0540U (effective 04/01/2025) | Transplantation medicine, quantification of donor?derived cell-free DNA using next-generation sequencing analysis of plasma, reported as percentage of donor?derived cell-free DNA to determine probability of rejection | |
| 0544U (effective 04/01/2025) | Nephrology (transplant monitoring), 48 variants by digital PCR, using cell-free DNA from plasma, donor derived cell-free DNA, percentage reported as risk for rejection | |
| HCPCS | No code | |
| ICD-10-CM (effective 10/01/15) | Investigational for all relevant diagnoses | |
| T86.00 | Unspecified complication of bone marrow transplant | |
| T86.01 | Bone marrow transplant rejection | |
| T86.02 | Bone marrow transplant failure | |
| T86.03 | Bone marrow transplant infection | |
| T86.09 | Other complications of bone marrow transplant | |
| T86.10 | Unspecified complication of kidney transplant | |
| T86.11 | Kidney transplant rejection | |
| T86.12 | Kidney transplant failure | |
| T86.13 | Kidney transplant infection | |
| T86.19 | Other complication of kidney transplant | |
| T86.20 | Unspecified complication of heart transplant | |
| T86.21 | Heart transplant rejection | |
| T86.22 | Heart transplant failure | |
| T86.23 | Heart transplant infection | |
| T86.290 | Cardiac allograft vasculopathy | |
| T86.298 | Other complications of heart transplant | |
| T86.30 | Unspecified complication of heart-lung transplant | |
| T86.31 | Heart-lung transplant rejection | |
| T86.32 | Heart-lung transplant failure | |
| T86.33 | Heart-lung transplant infection | |
| T86.39 | Other complications of heart-lung transplant | |
| T86.40 | Unspecified complication of liver transplant | |
| T86.41 | Liver transplant rejection | |
| T86.42 | Liver transplant failure | |
| T86.43 | Liver transplant infection | |
| T86.49 | Other complications of liver transplant | |
| T86.5 | Complications of stem cell transplant | |
| T86.810. | Lung transplant rejection | |
| T86.811 | Lung transplant failure | |
| T86.812 | Lung transplant infection | |
| T86.818 | Other complications of lung transplant | |
| T86.819 | Unspecified complication of lung transplant | |
| T86.820 | Skin graft (allograft) rejection | |
| T86.821 | Skin graft (allograft) (autograft) failure | |
| T86.822 | Skin graft (allograft) (autograft) infection | |
| T86.828 | Other complications of skin graft (allograft) (autograft) | |
| T86.829 | Unspecified complication of skin graft (allograft) (autograft) | |
| T86.830 | Bone graft rejection | |
| T86.831 | Bone graft failure | |
| T86.832 | Bone graft infection | |
| T86.838 | Other complications of bone graft | |
| T86.839 | Unspecified complication of bone graft | |
| T86.840 | Corneal transplant rejection | |
| T86.841 | Corneal transplant failure | |
| T86.842 | Corneal transplant infection | |
| T86.848 | Other complications of corneal transplant | |
| T86.849 | Unspecified complication of corneal transplant | |
| T86.850 | Intestine transplant rejection | |
| T86.851 | Intestine transplant failure | |
| T86.851 | Intestine transplant infection | |
| T86.852 | Other complications of intestine transplant | |
| T86.858 | Unspecified complication of intestine transplant | |
| T86.859 | Other transplanted tissue rejection | |
| T86.890 | Other transplanted tissue failure | |
| T86.891 | Other transplanted tissue infection | |
| T86.892 | Other transplanted tissue infection | |
| T86.898 | Other complications of other transplanted tissue | |
| T86.899 | Unspecified complication of other transplanted tissue | |
| T86.90 | Unspecified complication of unspecified transplanted organ and tissue | |
| T86.91 | Unspecified transplanted organ and tissue rejection | |
| T86.92 | Unspecified transplanted organ and tissue failure | |
| T86.93 | Unspecified transplanted organ and tissue infection | |
| T86.99 | Other complications of unspecified transplanted organ and tissue | |
| Z94.0 | Transplanted organ and tissue status, unspecified | |
| Z94.89 | Other transplanted organ and tissue status | |
| Z94.84 | Stem cells transplant status | |
| Z94.83 | Pancreas transplant status | |
| T86.20-T86.298 | Complications of heart transplant code range | |
| Z48.21 | Encounter for aftercare following heart transplant | |
| Z48.22 | Encounter for aftercare following kidney transplant | |
| Z94.0 | Kidney transplant status | |
| Z94.1 | Heart transplant status | |
| Z94.2 | Lung transplant status | |
| Z94.3 | Heart and lungs transplant status | |
| Z94.4 | Liver transplant status | |
| Z94.5 | Skin transplant status | |
| Z94.6 | Bone transplant status | |
| Z94.7 | Corneal transplant status | |
| Z94.81 | Bone marrow transplant status | |
| Z94.82 | Intestine transplant status | |
| ICD-10-PCS (effective 10/01/15) | Not applicable. ICD-10-PCS codes are only used for inpatient services. There are no ICD procedure codes for laboratory tests. |
Procedure and diagnosis codes on Medical Policy documents are included only as a general reference tool for each policy. They may not be all-inclusive.
This medical policy was developed through consideration of peer-reviewed medical literature generally recognized by the relevant medical community, U.S. FDA approval status, nationally accepted standards of medical practice and accepted standards of medical practice in this community and other nonaffiliated technology evaluation centers, reference to federal regulations, other plan medical policies, and accredited national guidelines.
"Current Procedural Terminology © American Medical Association. All Rights Reserved"
History From 2013 Forward
| 04/15/2025 | Annual review, updating frequency of testing in criteria #3 one test per month at one/two/three/four months post transplant and one test every three months beginning at six months post transplant. Also updating description, rationale, guidelines/recommendations, regulatory status , and references. |
| 02/11/2025 | Updated coding section. Added codes 0540U and 0544U. These codes will be effective 04/01/2025. No other changes. |
| 01/22/2025 | Added CPT code 81558 effective 01/01/2025 |
| 09/05/2024 | Updated CPT coding. Added codes 0493U, 0508U and 0509U (effective 10/01/2024). No change in policy intent. |
| 09/04/2024 | Updating coding. Updated verbiage on Code 0118U (effective date 10/01/2024) |
| 08/01/2024 | Annual review, no change to policy intent. Updating description, rationale, references. |
| 01/25/2024 | Annual review, updating policy to include coverage criteria for AlloSure Heart, and Prospera Heart. Adding a not medically necessary statement for HearetCare. Updating table of terminology, rationale and references. |
| 11/06/203 | Annual review, entire policy updated for clarity and consistency. Adding criteria #6 to assess for rejection and injury using mRNA expression profiling as not medically necessary. |
| 11/07/2022 | Annual review, no change to policy intent. Updating policy for clarity. Also updating rationale, references, and coding |
| 04/12/2022 |
Interim review to update coding: 0319U, 0320U, 84999 (for Heartsbreath), removing 0018M. NO other changes made. |
| 10/01/2021 |
Annual review, updating policy to include coverage statement regarding TruGraf. Also updating coding, background, rationale and references. |
| 10/01/2020 |
Annual review. No change to policy intent. |
| 10/22/2019 |
Annual review with extensive revision to policy. Expanding policy from heart transplant rejection to transplant rejection testing. Policy verbiage to expand testing in non heart related transplant testing. Reformatting policy for clarity. |
| 10/29/2018 |
Annual review, no change to policy intent. |
| 11/02/2017 |
Annual review, rewriting policy for clarity and specificity. No change to policy intent. |
| 04/26/2017 |
Updated category to Laboratory. No other changes. |
| 11/07/2016 |
Annual review, updating policy verbiage for clarity. |
| 01/05/2016 |
Udated cpt code. No other changes made to policy |
| 12/14/2015 |
Annual review, changed verbiage to allow Allomap testing as medically necessary. Also updated guidelines, background, rationale, references and coding. |
| 12/1/2015 |
Updated CPT codes with 2016 codes. No change policiy intent. |
| 01/13/2015 |
Annual review, no change to policy intent. Updated background, rationale, and references. Added guidelines and coding. |
| 01/14/2014 |
Annual review. Added benefit application & related policies. Updated rationale and references. No change to policy intent. |